The genome of *Mediocremonas mediterraneus*
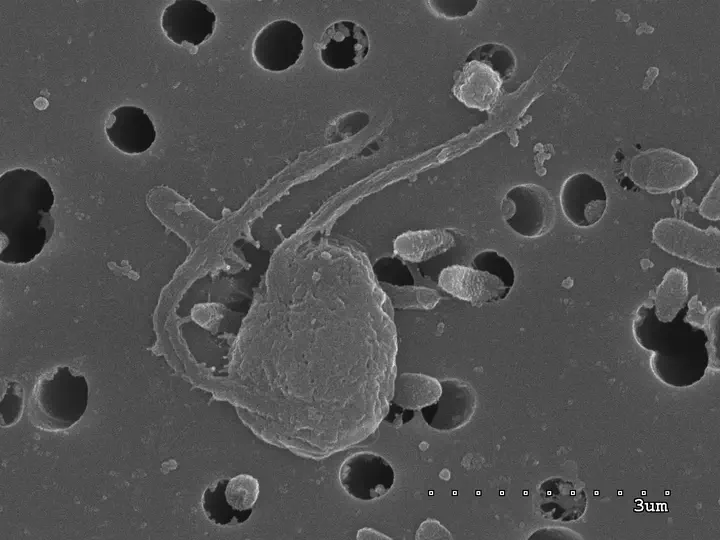 Photo by Javier del Campo
Photo by Javier del CampoThere is a significant bias in eukaryotic genomics that impoverishes our understanding of eukaryotic diversity: most genomics research focuses on multicellular eukaryotes. Around 85% of sequenced eukaryotic genomes correspond to multicellular organisms — Metazoa, Fungi, or Land Plants — yet these lineages represent only ~23% of all operational taxonomic units (OTUs) in environmental surveys. This leaves the vast majority of eukaryotic diversity underrepresented in genomic databases, skewing our views of what a eukaryote is and what roles they play in the environment.
In this project, carried out as part of the Catalan Initiative for the Earth Biogenome Project, we are generating the reference genome of Mediocremonas mediterraneus, a heterotrophic nanoflagellate isolated from Blanes Bay (Catalonia). Mediocremonas mediterraneus belongs to the Developea within the supergroup Stramenopiles. Based on phylogenomics, the Developea are sister to all photosynthetic Stramenopiles (diatoms, kelps, etc.), making M. mediterraneus an ideal candidate to study the evolutionary origins of photosynthesis in Stramenopiles.